Neural Network Application for Alzheimer's Disease Identification Using EEG Data
University of Toronto — APS360 Applied Fundamentals of Deep Learning
Garvish Bhutani, Radhika Banerjea, Sunny Zhang
Abstract
This project investigates the use of a deep convolutional neural network to identify patients with Alzheimer's disease (AD), Frontotemporal Dementia (FTD), or healthy controls using 19-channel EEG data. Current diagnostics only detect Alzheimer's in later stages — our goal was to leverage the subtle neurological changes visible in EEG signals for earlier detection. The final 5-layer 1D CNN outperformed all existing models on this dataset, achieving 70% test accuracy and 100% recall for Alzheimer's patients, compared to a 44% baseline from a graph signal processing approach.
Technologies
Problem & Motivation
Alzheimer's is the most common form of dementia, causing progressive brain cell death with no known cure. Current diagnostics only detect it in later stages when symptoms are already severe — limiting treatment options and quality of care.
Because Alzheimer's is neurological, EEG signals should encode early biomarkers before symptoms appear. However, unlike epilepsy (which has sharp, distinguishable seizure signals), Alzheimer's manifests as subtle shifts in specific frequency bands — making it particularly hard for supervised models to latch onto, and nearly impossible to hand-engineer features for.
This made deep learning the natural approach: automatic feature extraction without requiring prior knowledge of what to look for.
Data Processing
The dataset (OpenNeuro ds004504) contains 88 subjects with 19 channels of EEG data averaged over 13.2 minutes, stored in BIDS-format `.set` files and parsed with the MNE Python library.
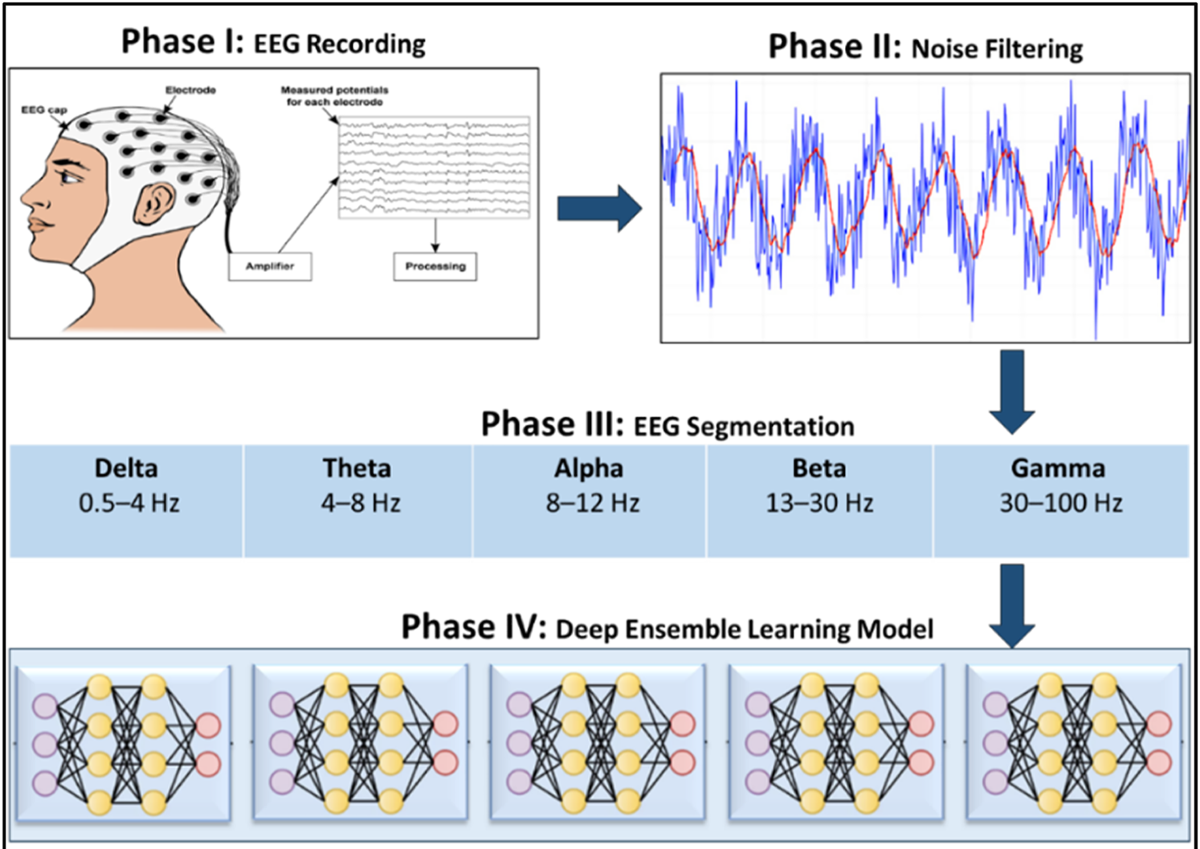
Bandpass Filtering: A Butterworth bandpass filter (0.5–45 Hz) was applied to remove out-of-band noise. These cutoffs match those used by the dataset creators, enabling comparison between our preprocessing and theirs.
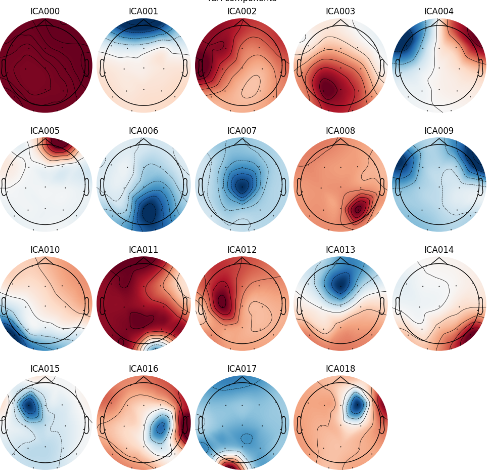
Artifact Removal: Independent Component Analysis (ICA) isolated and removed signal components corresponding to eye blinks, eye movements, line noise, and heartbeat artifacts — standard practice in EEG analysis.
We preprocessed the raw data ourselves rather than using the provided preprocessed version, which had discontinuities that could interfere with training. Our preprocessing closely matched the provided version without those discontinuities.
Model Architecture
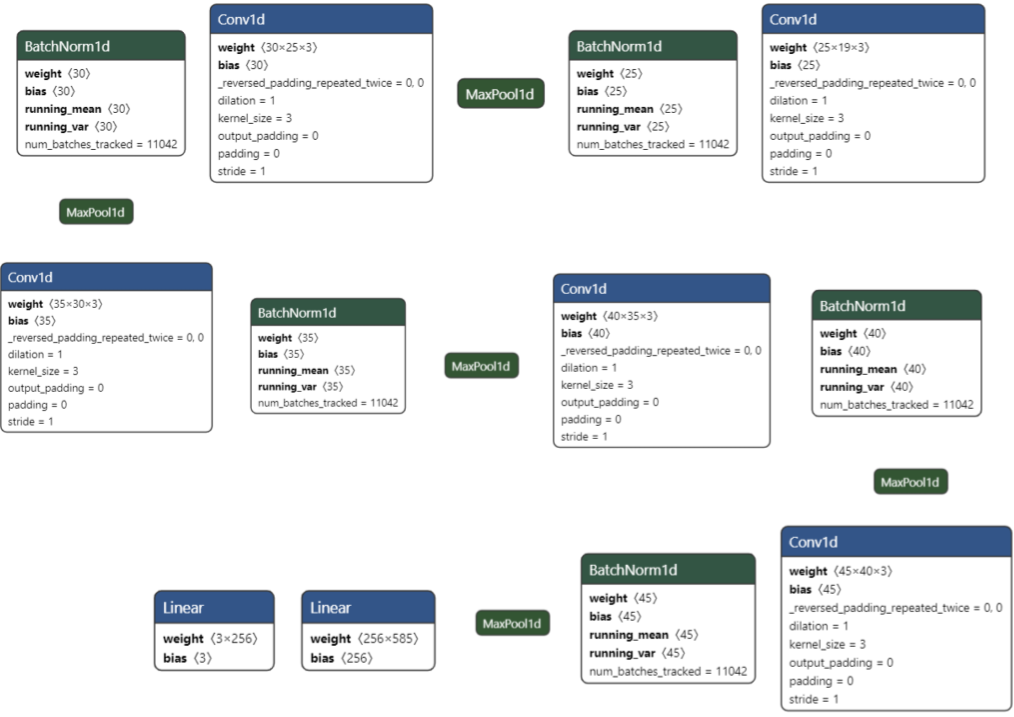

After evaluating 50+ architectures, the final model is a 5-layer 1D CNN that progressively upsamples the EEG channel dimension:
19 → 25 → 30 → 35 → 40 → 50 channels across 5 Conv1D layers, each followed by BatchNorm1d and MaxPool1d (kernel=4, stride=4).
Key design choices validated through ablation: - Upsampling outperforms downsampling for this signal structure - MaxPool > AvgPool, and larger kernel/stride (4,4) beats (2,2) - ReLU > Leaky ReLU across all tested configurations - SGD > Adam — Adam consistently underperformed on both 3-layer and 5-layer variants - Exponential LR decay outperformed constant and linear schedules - 2 linear layers map the final features to 3 classes (Alzheimer's, FTD, Healthy) with Softmax output
LSTMs were evaluated but ruled out — they require well-encoded temporal features (present in epilepsy), but Alzheimer's lacks the acute signal events needed to make LSTMs effective.
Results
The final model achieved 70% test accuracy on 40 held-out subjects — 26% higher than the best baseline (a graph signal processing approach using Naive Bayes at 44%) and more than double the simple ANN baseline (33%).
Per-class performance: - Healthy: 62.5% precision, 100% recall, 76.9% F1 - Alzheimer's: 76.9% precision, 100% recall, 86.9% F1 - FTD: 100% precision, 20% recall, 33.3% F1
The model excels at identifying Alzheimer's and Healthy patients, but struggles with FTD — likely because FTD's EEG signature overlaps heavily with healthy signals, and the dataset had fewer FTD samples. The model tends to classify FTD patients as healthy, which is an important limitation for clinical use.
High variance across training runs (sometimes spanning 50% test accuracy range) suggests the model occasionally latches onto spurious features, likely due to the subtle and inconsistent nature of Alzheimer's EEG signatures.
Ethical Considerations
Diagnostic transparency: Reporting test accuracy alone is insufficient. The model under-diagnoses FTD — misclassifying it as healthy — which would be clinically dangerous if deployed without this context clearly communicated.
Dataset bias: With only 88 subjects, the model cannot be comprehensively evaluated for racial, gender, or age-based biases. Any clinical deployment would require substantially larger and more diverse datasets.
The model should be understood as a research prototype demonstrating the feasibility of deep learning for EEG-based Alzheimer's detection, not a production diagnostic tool.